A foundation model for breast and lung cancer screening using non-contrast computed tomography
Ethics approval
All retrospective non-public datasets (Sites A, B and G) in this investigation were approved by the institutional review board (IRB) of the hospitals with a waiver granted for the requirement of informed consent. With respect to the prospective study pre-registered at www.chictr.org.cn (identifier ChiCTR2400081249), all participants signed an informed consent developed and approved by the IRB of Sites C, D, E and F. All datasets were de-identified before model development and test in this investigation.
Chest CT dataset
Our study incorporated ten distinct CT datasets, including six Chinese (Sites A to F) and four international public datasets (CT-RATE, NLST, LIDC and LungCT). These datasets represented diverse clinical settings (emergency rooms, physical examination centres, inpatient and outpatient departments) and included scans from seven manufacturers (GE, Philips, SIEMENS, TOSHIBA, MinFound, UIH and Neusoft). Site A, Site B and all public datasets were characterized as retrospective cohorts used for the development and testing of the OMAFound model, while the remaining datasets (Sites C to F) provided prospective low-dose CT scans from screening populations for real-world validation.
The datasets were categorized into two types based on clinical interpretation availability. The first type consisted of unlabelled data (Site A-CTunlabeled and CT-RATE), which provided large-scale datasets exclusively for task-agnostic foundation model pretraining. The second type was weakly supervised labelled data with patient-level ground-truth status, confirmed either by pathology (cancer or non-cancer) or at least 2 years (unless otherwise specified) follow-up for non-cancer status confirmation. Within the labelled data, two labelling patterns emerged: retrospective datasets (Site A-CTbreast, Site A-CTlung, Site B, NLST, LIDC and LungCT) contained a single label per patient (either breast or lung), while prospective datasets (Sites C to F) provided comprehensive dual labelling, including both breast and lung assessments for each patient.
For model training, all eligible examinations per patient were utilized, whereas only a single CT scan per patient was used for model testing. To prevent the risk of label leakage, anonymized patient IDs were used across all datasets, ensuring no patient overlaps between training and test cohorts (all scans from the same patient were assigned to the same cohort). Table 1 and Extended Data Fig. 1 provide comprehensive details on dataset utilization and patient assignment criteria. Additional dataset specifications are provided below.
Site A (The First Affiliated Hospital of Anhui Medical University). Data were retrospectively collected from multiple clinical settings (emergency rooms, inpatient and outpatient departments) between October 2015 and April 2024, which were subsequently divided into unlabelled and labelled datasets. The Site A-CTunlabeled dataset comprised 159,273 unlabelled CT scans from 37,507 patients. The labelled data were further categorized into Site A-CTbreast dataset, containing scans from 16,007 non-cancer patients and 6,754 patients with breast cancer, and Site A-CTlung dataset, consisting of scans from 23,785 non-cancer patients and 3,672 patients with lung cancer. For the organ-specific adaptation phase, labelled data were randomly and selectively allocated to the fine-tuning cohort (most cancer cases were used here to alleviate class imbalance issue on training) and the internal test cohort.
Site B (No.2 People’s Hospital of Fuyang City). Standard-dose CT scans were retrospectively collected from the outpatient department between February 2020 and May 2024, resulting in a total of 1,716 labelled CT from 1,716 patients (1,661 non-cancer patients and patients with 55 breast cancer). Site B was used solely for external testing of the breast module of OMAFound to assess generalizability.
Site C (physical examination centres affiliated to Site A). Low-dose CT scans were collected through a pre-registered prospective study. A total of 10,680 screening participants were enroled between January 2024 and December 2024. The cohort comprised 10,603 non-cancer cases, confirmed through 6–12 months of short-term follow-up. The remaining cases included 15 breast cancer cases and 62 lung cancer cases (24 from female and 38 from male), all confirmed by pathology results. Site C was used solely for prospective real-world assessment of OMAFound in multi-cancer screening.
Site D (Lu’an People’s Hospital). Low-dose CT scans were prospectively collected from 1,214 screening participants between January 2024 and July 2024. Disease statuses were determined through either 6–12 months of short-term follow-up or pathology confirmation, identifying 1,192 non-cancer cases and 22 cancer cases (12 breast cancer, 4 female lung cancer and 6 male lung cancer). Site D was used solely for prospective real-world assessment of OMAFound in multi-cancer screening.
Site E (Weifang Traditional Chinese Hospital). Between January 2024 and December 2024, a total of 5,181 low-dose CT scans were prospectively collected during annual physical examinations. These scans represented 5,140 non-cancer patients, 14 patients with breast cancer and 27 patients with lung cancer (14 from female and 13 from male). Site E was used solely for prospective real-world assessment of OMAFound in multi-cancer screening.
Site F (Xuancheng People’s Hospital). We prospectively enroled participants from a local screening population for low-dose CT scans. Following standardized prospective labelling criteria, 4,426 non-cancer patients, 43 patients with breast cancer and 57 patients with lung cancer (35 from female and 22 from male) were finally collected between January 2024 and December 2024. Site F was used solely for prospective real-world assessment of OMAFound in multi-cancer screening.
CT-RATE (non-contrast chest CT dataset32). This public dataset was collected at Istanbul Medipol University Mega Hospital between May 2015 and January 2023. It comprises 50,188 unlabelled CT data from 21,304 unique patients. CT-RATE was used solely for task-agnostic foundation model pretraining.
NLST (National Lung Screening Trial42). The NLST dataset was collected across 33 US medical institutions, with participants randomized to receive annual low-dose CT screenings between August 2002 and 2007. In total, 41,805 labelled CT scans from 19,698 patients (18,717 non-cancer patients and 981 patients with lung cancer) were included, with long-term follow-up data available. A random subset (12.7%) at the patient level was allocated to the internal test cohort, while the remaining scans were used for training. NLST was used solely for multi-year lung cancer risk prediction, where a single low-dose CT scan was used to predict lung cancer occurrence 1–6 years post-screening.
PublicX (combined LIDC40 and LungCT41 datasets). The LIDC dataset with a mix of standard-dose and low-dose scans were collected from five different institutions between 1998 and 2010. The LungCT dataset contains standard-dose CT scans acquired between July 2004 and June 2011. On the basis of the same inclusion criteria for the nationwide dataset, the PublicX dataset includes 396 labelled CT data from 396 patients (162 non-cancer patients and 234 patients with lung cancer). The PublicX dataset was used solely for external testing of the lung module of OMAFound to assess generalizability.
Mammography dataset
Given mammography’s status as the current gold standard for breast cancer screening, we developed a mammography-based AI model as a benchmark for comparison with the CT-based OMAFound. For this purpose, we retrospectively collected a dedicated mammography-only dataset, designated as Site A-MG to distinguish it from chest CT data of Site A, for the development and evaluation of this mammography-based AI model.
Specifically, Site A-MG includes 72,116 mammography images from 18,029 patients (bilateral cranial–caudal and mediolateral oblique views per patient), acquired between January 2014 and December 2023 from either a GE Senographe DS mammography system or Hologic Selenia Dimensions mammography system, covering both screening and diagnostic populations. To assess the generalizability of our mammography-based AI model, we assembled an external test cohort from Anhui No.2 Provincial People’s Hospital (Site G). This cohort contained 3,280 mammography images from 820 patients (158 cancer-positive cases), retrospectively collected between March 2023 and August 2024 using a GE Senographe DS mammography system.
The labels of these mammography datasets were confirmed either by pathology (cancer or non-cancer) or through a minimum follow-up period of 2 years for non-cancer status confirmation. Detailed patient characteristics and labels are provided in Extended Data Table 1.
Paired CT–mammography dataset
Recognizing that model performance can vary across different populations and clinical settings, we thus established a more equitable comparison between the mammography-based AI model and CT-based OMAFound for breast cancer screening. That is, we additionally collected 1,131 paired CT and mammography scans from 1,131 patients (Extended Data Table 1), namely, as Site A-CTMG. Importantly, Site A-CTMG data had no overlap with either Site A-CTbreast or Site A-MG datasets.
OMAFound model
Image preprocessing before OMAFound model development was performed using Torchvision (version 0.20.1) and SciPy (version 1.14.1). The multi-institutional CT dataset showed slice spacing variations from 0.625 mm to 5 mm. To harmonize the difference in slice thickness and spatial resolution, all CT scans were resampled to a uniform 1 × 1 × 1 mm before resizing to voxel dimensions of 128 × 128 × 128. Intensity distributions (Hounsfield units) were standardized using min–max normalization, and foreground regions of lung window and soft tissue window were extracted from each scan. In this study, the model development process did not incorporate any image annotations, such as lesion bounding boxes or segmentation masks.
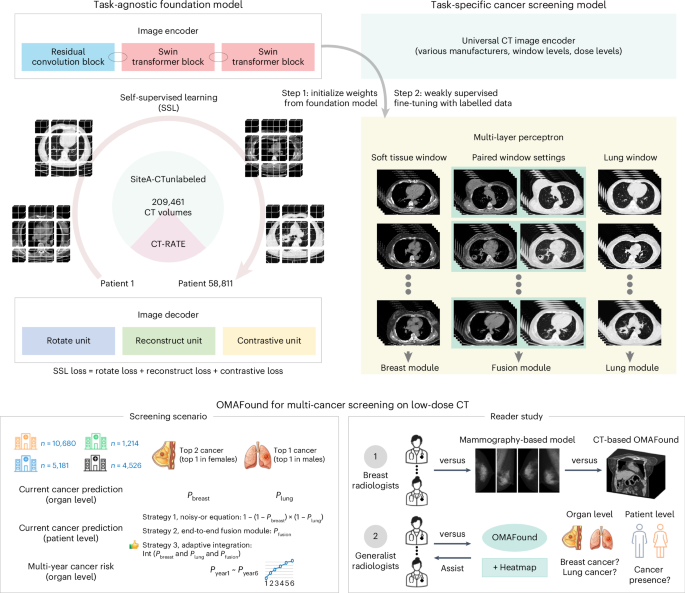
The architecture of the SSL-based OMAFound model is detailed in Supplementary Fig. 1 and the task-specific downstream modules are shown in Supplementary Fig. 2. For the foundation model, we used the encoder from SwinUNETR-V235 as the backbone for feature extraction, integrating 3D stage-wise convolution and shifted window-based self-attention mechanisms. A residual convolution (ResConv) block was added at the beginning of each resolution level, followed by a Swin transformer block.
In the organ-specific breast and lung modules, a 3D adaptive average pooling layer was utilized to aggregate spatial features, followed by a fully connected layer and softmax activation for cancer risk prediction task. Specifically, the breast module and lung module of OMAFound were developed using the fine-tuning cohort of Site A-CTbreast and Site A-CTlung, respectively.
For the fusion module, the encoders for the breast and lung branches were initialized with weights from the corresponding organ-specific modules and kept frozen during fusion training. Each encoder produced a 768-dimensional feature vector, which was used to generate classification logits and uncertainty estimates. A learnable class token was concatenated with the two feature vectors and passed through a transformer encoder to capture cross-organ interactions. The final cancer prediction was derived from the updated class token, and the total loss was calculated as the sum of the fusion loss and organ-specific uncertainty losses. The fusion module was developed using combined fine-tuning datasets from both breast and lung modules and tested on merged internal test cohorts of Site A-CTbreast and Site A-CTlung.
OMAFound was implemented using the PyTorch framework (version 2.5.1), and training was conducted using two Intel Xeon central processing units and eight NVIDIA A100 80GB graphics processing units. Inspired by previous research52, the objective of the SSL module was to minimize a combination of rotation loss, reconstruction loss and contrastive loss. For downstream tasks, label smoothing loss was applied. Optimization was performed using the adaptive moment estimation (ADAMW) optimizer, with a batch size of 96 and an initial learning rate of 0.0001. A linear warm-up ratio of 0.1 was applied, followed by a cosine function learning rate schedule. Training was capped at 15 epochs, with early stopping triggered if no further loss improvement was observed.
To address class imbalance, weighted sampling was used to ensure balanced representation of all classes during training. Data augmentation included random affine transformations (translation and scaling within the bounds of (0.1, 0.1, 0.1), random rotations (up to 15°), contrast adjustment with a random factor between 0.8 and 1.2, and the addition of random noise with intensities ranging from 0.005 to 0.05. All augmentations were constrained to maintain pixel values within the [0, 1] range.
Mammography-based AI model
To compare chest CT with the standard mammography-based approach for breast cancer screening, we developed an individual mammography-based AI model using the dataset from Site A-MG. Mammography scans containing both cranial–caudal and mediolateral oblique views of the bilateral breast were included for model development.
Supplementary Fig. 3 illustrates the architecture of the mammography-based AI model. The model, a derivative of BMU-Net20, integrates a ResNet-18 backbone with a transformer encoder for multi-view breast cancer classification. The ResNet-18 backbone, initialized with weights transferred from the large-scale, pre-trained Mirai model21, was used to extract features from each individual view. These features were then augmented with positional embeddings and passed through the transformer encoder to capture contextual dependencies across views. Separate classifiers were applied to each view, and their outputs were weighted by learnable parameters specific to the left and right sides. The final logit was obtained by averaging the weighted outputs.
Reader study on mammography
We conducted a mammography reader study to compare the performance of the mammography-based AI model with that of experienced breast radiologists. To be specific, each reader independently reviewed the same set of cases and assigned a BI-RADS (Breast Imaging Reporting and Data System) 5th edition53 rating using the values 1, 2, 3, 4a, 4b, 4c and 5, simulating routine clinical interpretation. To convert BI-RADS assessments into binary classification for sensitivity and specificity calculations, BI-RADS 4a or higher were considered as test positive, and all others negative. The average reader sensitivity and specificity were computed by averaging the individual sensitivity and specificity values across all readers. All readers were blinded to each other’s assessments, the original clinical reports and the AI model outputs. The study included 5 board-certified radiologists specializing in mammography, each with over 10 years of clinical experience. A total of 190 examinations—randomly selected from the test cohort of the Site A-CTMG dataset—were presented to the readers in a randomized order.
Reader study on low-dose CT
To evaluate the clinical utility of OMAFound in assisting generalist radiologists with improved screening outcomes, we conducted a 2-part CT reader study involving 365 patients (220 non-cancer, 59 breast cancer, 34 female lung cancer and 52 male lung cancer). Cases were randomly and selectively sampled (higher for cancer cases and lower for non-cancer cases to enhance the difficulty of the screening task and statistical power) from the prospective cohorts of Sites C, D, E and F. Seven board-certified generalist radiologists participated in this study, with their clinical experience summarized in Extended Data Table 4.
The sequential reader study consisted of a first reading (solo) and a second reading (+OMAFound). Each reader was requested to finish three tasks, including organ-level breast cancer detection, organ-level lung cancer detection and patient-level cancer presence prediction. During the first reading, each reader independently reviewed the same set of testing cases without time limit and provided initial binary decisions for each task (‘Yes’ for cancer, ‘No’ for non-cancer). In the second reading, readers were provided with OMAFound-generated heatmaps and prediction scores as a decision support. They were allowed to update their initial assessments based on the AI assistance.
Interpretability of the OMAFound model
To assure trust from human experts, it is essential to make the model’s decision-making process interpretable. In this study, we implemented and analysed five post-hoc explanation approaches, including four CAM-based (Grad-CAM54, Grad-CAM + + 55, Layer-CAM56 and Finer-CAM57) and one attention-based gradient-driven multi-head attention rollout (GMAR58) mappings, to visualize the heatmap localization regions that could aid human experts to understand the justification of the AI system for the cancer risk predictions. All post hoc methods in this study were applied to the normalization layer of the final stage of the model for each test image.
Specifically, Grad-CAM++ enhances Grad-CAM by implementing pixel-wise weights instead of channel-wise weights, improving small object localization capability. Layer-CAM generates more reliable boundary definitions by utilizing pixel-level activation with positive gradients within and across layers. Finer-CAM extends Layer-CAM by incorporating progressive cross-layer refinement and denoising, achieving superior semantic alignment. GMAR is a novel method to quantify the importance of each attention head using gradient-based scores.
Statistical analysis
The performance of the OMAFound model and the mammography-based AI model was evaluated using weighted F1 score, balanced accuracy, sensitivity, specificity and the AUC. The 95% CIs of the weighted F1 score, balanced accuracy and specificity were computed using 1,000 non-parametric bootstrap resamples. A dynamic approach (Wilson CIs and bootstrap-based CIs) was used for sensitivity due to low cancer prevalence. The C-index43 was computed to evaluate the predictive performance of time-to-event models. AUC comparisons were conducted using Delong’s test. All comparisons were two-sided, with a P value <0.05 considered statistically significant. All statistical analyses were performed using SPSS (version 22.0), and relevant Python packages.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
link